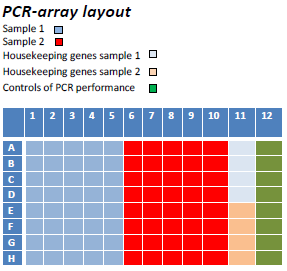
Array layout, Sea bream Mito-Chip
Array of selected markers of mitochondrial biogenesis and activity
PCR-ArraysBermejo-Nogales et al., 2014, General and Comparative Endocrinology 205; 305-315
Array layout, Sea bream OXPHOS-Chip
A complete set of oxidative phosphorylation (88 genes)-mitochondrial respiratory chain, including mitochondrial- and nuclear-encoded genes of complexes I, II, III, IV and V, and their assembly proteins
PCR-ArraysArray layout, Sea bream Lipid-Chip
Array of 40 selected markers of lipid metabolism
PCR-ArraysArray layout, Sea bream Interleukin-Chip
Array of 19 immunorelevant genes, including ILs and IL-receptors
PCR-ArraysPérez-Cordón et al., 2014, Fish & Shellfish Immunology 37; 201-208
Sea bream- Circadian gene regulation in whole larva
Customized Agilent oligoarray (8x15 k); 13939 annotated genes (sea_bream_nutrigroup_array v.3)
MicroarraysArray layout, Sea bream Oxidative Stress-chip
Array of 34 selected markers of oxidative stress
PCR-ArraysPérez-Sánchez et al., 2013 Comparative Biochemistry & Physiology 8D; 123-130
Sea bass- Funtional specialization along the intestinal transcriptome
Customized Agilent oligoarray (8x15 k); 14147 different annotated genes (sea_bass_nutrigroup_array v.1)
MicroarraysArray layout, Sea bream Gut-chip
Array of 88 selected markers of intestinal function and integrity.
PCR-ArraysPérez-Sánchez et al., 2015. Fish & Shellfish Immunology 44, 117-128
Estensoro et al., 2016. PLoS ONE 11:e0166564
Gisbert et al., 2017. British Journal of Nutrition 117, 351-363
Sea bream- Tissue-specific response after hyper- and hypo-osmotic challenges
Customized Agilent oligoarray (4 x 40 k); 6862 annotarted sequences + 8294 non-annotated sequences (AQUAGENOMICS array)
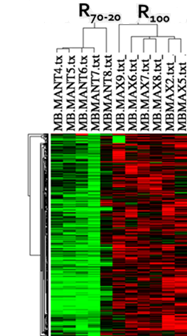
MicroarraysMuscle transcriptome/Meta-analysis of microarray gene expression profiling in fish
Customized Agilent oligoarray (4 x 40 k); 6862 annotarted sequences + 8294 non-annotated sequences (AQUAGENOMICS array)
Meta-analysis tool (Fish & Chips, AQUAEXCEL Poject)
MicroarraysSea bream- Nutritional regulation of intestinal transcriptome
Customized Agilent oligoarray (8x15 k); 7000 annotated genes genes (AQUAMAX array, sea_bream_nutrigroup_array v.2)
MicroarraysSea bass, Lipid-Chip
Array of 29 selected markers of lipid metabolism
PCR-ArraysRimoldi et al., 2016. Reviews in Fish Biology and Fisheries 26, 93-108
Sea bream- Gene expression profiling of head kidney and intestine in parasitized fish
18490 cDNA clones (AQUAFIRST array, sea_bream_nutrigroup_array v.1)
MicroarraysSea bream- Gene expression profiling of liver in crowded stressed fish
18490 cDNA clones (AQUAFIRST array, sea_bream_nutrigroup_array v.1)
MicroarraysSea bream, Growth-Chip
Array of 87 selected growth markers, including markers of GH/IGF system, muscle growth and cell differentiation and proliferation, protein folding, assembly and breakdown, and inflammatory or anti-inflammatory responses.
PCR-Arrays