MICROARRAYS
Sea bream- Circadian gene regulation in whole larva
Customized Agilent oligoarray (8x15 k); 13939 annotated genes (sea_bream_nutrigroup_array v.3)
MicroarraysSea bass- Funtional specialization along the intestinal transcriptome
Customized Agilent oligoarray (8x15 k); 14147 different annotated genes (sea_bass_nutrigroup_array v.1)
MicroarraysSea bream- Tissue-specific response after hyper- and hypo-osmotic challenges
Customized Agilent oligoarray (4 x 40 k); 6862 annotarted sequences + 8294 non-annotated sequences (AQUAGENOMICS array)
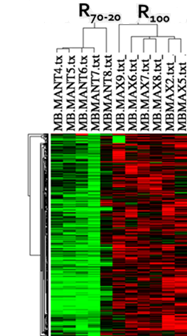
MicroarraysMuscle transcriptome/Meta-analysis of microarray gene expression profiling in fish
Customized Agilent oligoarray (4 x 40 k); 6862 annotarted sequences + 8294 non-annotated sequences (AQUAGENOMICS array)
Meta-analysis tool (Fish & Chips, AQUAEXCEL Poject)
MicroarraysSea bream- Nutritional regulation of intestinal transcriptome
Customized Agilent oligoarray (8x15 k); 7000 annotated genes genes (AQUAMAX array, sea_bream_nutrigroup_array v.2)
MicroarraysSea bream- Gene expression profiling of head kidney and intestine in parasitized fish
18490 cDNA clones (AQUAFIRST array, sea_bream_nutrigroup_array v.1)
MicroarraysSea bream- Gene expression profiling of liver in crowded stressed fish
18490 cDNA clones (AQUAFIRST array, sea_bream_nutrigroup_array v.1)
Microarrays